May 21 2022
... arambaut
Some thoughts on the long branch to 2022 outbreak genomes and APOBEC3 editing:
It seems to me that APOBEC3 has caused much of the single nucleotide variability in this MPXV clade (I have yet to investigate the other clades). These enzymes act as anti-viral defences in mammals acting on single stranded DNA and deaminating cytosine to create uracil bases which then get paired with an adenine base as the other strand is synthesised resulting in a G → A mutation. Given there is the signal of editing on both strands, my guess is that this will happen in a single cell during multiple rounds of genome replication.
Presumably this happens a lot but in most cases such intensive random mutation will induce changes that will destroy the function of a protein rendering the virus inactive. However, occasionally a virus will not be sufficiently hit by APOBEC3 to deactivate it and it will continue to replicate and transmit. Whilst the branch leading to the 2022 outbreak is exceptionally long, under this model we would predict that all or most of these mutations arose in a single round of replication. Thus under the APOBEC3 mutation model would be that we will not see ‘intermediate’ genomes with only some of the 41 mutations. Although the 4 year gap in genome sequencing could mean that the long branch is the result of more than one bout of APOBEC3 editing in different hosts.
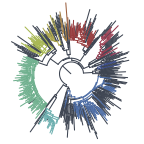
Given there are other branches in this tree with similar patterns of APOBEC3-like mutation, I would infer that this occurred in the reservoir host (possibly rodent species). I expect that on the short term, all further genomes from the current outbreak in Europe will be essentially identical. Cases that are not part of the outbreak – possibly representing independent emergence or other outbreaks – may not have these mutations. Alternatively this variant may have become the dominant variant in the reservoir (similar to that seen in for the 2018 clade from which the 2022 clade arose).
These mutations may allow a cheap and easy PCR-based genotyping system to be developed to track outbreaks rather than full-genome metagenomics (which will give diminishing returns given the normally low mutation rate and long genome).
gustavo_palacios
@arambaut Andrew, I think that the pattern of mutation that we are observing is puzzling, and deserves attention regarding what it means. The dinucleotide and tetra nucleotide patterns that you highlighted are solid. However, do you recollect other instances of APOBEC3 induced biased hypermutation that are so spread?
Most of the examples that I can recall are T-to-C and A-to-G changes in localized areas of the genome, and not spread over distant region of the genome as observed here.
I did not have the time to dig in deep in the literature enough, though, but all the examples in nature that I find appear to follow the pattern of biased hypermutation over small stretches of the genome. That also makes sense considering the mechanism of action of the cytidine-deaminase and the need of the DNA to be in a single strand state for action.
Have you find other examples like this where the APOBEC3 effects are spread over a 150kb region?
Puzzling. ...
... arambaut
Some thoughts on the long branch to 2022 outbreak genomes and APOBEC3 editing:
It seems to me that APOBEC3 has caused much of the single nucleotide variability in this MPXV clade (I have yet to investigate the other clades). These enzymes act as anti-viral defences in mammals acting on single stranded DNA and deaminating cytosine to create uracil bases which then get paired with an adenine base as the other strand is synthesised resulting in a G → A mutation. Given there is the signal of editing on both strands, my guess is that this will happen in a single cell during multiple rounds of genome replication.
Presumably this happens a lot but in most cases such intensive random mutation will induce changes that will destroy the function of a protein rendering the virus inactive. However, occasionally a virus will not be sufficiently hit by APOBEC3 to deactivate it and it will continue to replicate and transmit. Whilst the branch leading to the 2022 outbreak is exceptionally long, under this model we would predict that all or most of these mutations arose in a single round of replication. Thus under the APOBEC3 mutation model would be that we will not see ‘intermediate’ genomes with only some of the 41 mutations. Although the 4 year gap in genome sequencing could mean that the long branch is the result of more than one bout of APOBEC3 editing in different hosts.
Given there are other branches in this tree with similar patterns of APOBEC3-like mutation, I would infer that this occurred in the reservoir host (possibly rodent species). I expect that on the short term, all further genomes from the current outbreak in Europe will be essentially identical. Cases that are not part of the outbreak – possibly representing independent emergence or other outbreaks – may not have these mutations. Alternatively this variant may have become the dominant variant in the reservoir (similar to that seen in for the 2018 clade from which the 2022 clade arose).
These mutations may allow a cheap and easy PCR-based genotyping system to be developed to track outbreaks rather than full-genome metagenomics (which will give diminishing returns given the normally low mutation rate and long genome).
gustavo_palacios
@arambaut Andrew, I think that the pattern of mutation that we are observing is puzzling, and deserves attention regarding what it means. The dinucleotide and tetra nucleotide patterns that you highlighted are solid. However, do you recollect other instances of APOBEC3 induced biased hypermutation that are so spread?
Most of the examples that I can recall are T-to-C and A-to-G changes in localized areas of the genome, and not spread over distant region of the genome as observed here.
I did not have the time to dig in deep in the literature enough, though, but all the examples in nature that I find appear to follow the pattern of biased hypermutation over small stretches of the genome. That also makes sense considering the mechanism of action of the cytidine-deaminase and the need of the DNA to be in a single strand state for action.
Have you find other examples like this where the APOBEC3 effects are spread over a 150kb region?
Puzzling. ...



Comment